
Section Abstract Introduction Methods Results Discussion Conflict of Interest Acknowledgment References
Basic Medical Research
TMEPAI genome editing in triple negative breast cancer cells
pISSN: 0853-1773 • eISSN: 2252-8083
http://dx.doi.org/10.13181/mji.v26i1.1871 Med J Indones. 2017;26:14–8
Received: February 15, 2017
Accepted: April 06, 2017
Author affiliation:
1 Doctoral Program in Biomedicine, Faculty of Medicine, Universitas Indonesia, Jakarta, Indonesia
2 Medical Sciences Master Program, Graduate School of Comprehensive Human Sciences, University of Tsukuba, Ibaraki, Japan
3 Department of Experimental Pathology, Faculty of Medicine, University of Tsukuba, Ibaraki, Japan
4 Department of Pharmacology and Therapeutics, Faculty of Medicine, Universitas Indonesia, Jakarta, Indonesia
Corresponding author:
Yukihide Watanabe
E-mail: y-watanabe@md.tsukuba.ac.jp
Background
Clustered regularly interspaced short palindromic repeats/CRISPR-associated 9 (CRISPR/Cas9) is a powerful genome editing technique. It consists of RNA-guided DNA endonuclease Cas9 and single guide RNA (gRNA). By combining their expressions, high efficiency cleavage of the target gene can be achieved, leading to the formation of DNA double-strand break (DSB) at the genomic locus of interest which will be repaired via NHEJ (non-homologous end joining) or HDR (homology-directed repair) and mediate DNA alteration. We aimed to apply the CRISPR/Cas9 technique to knock-out the transmembrane prostate androgen-induced protein (TMEPAI) gene in the triple negative breast cancer cell line.
Methods
Designed gRNA which targets the TMEPAI gene was synthesized, annealed, and cloned into gRNA expression vector. It was co-transfected into the TNBC cell line using polyethylenimine (PEI) together with Cas9-GFP and puromycin resistant gene vector. At 24-hours posttransfection, cells were selected by puromycin for 3 days before they were cloned. Selected knock-out clones were subsequently checked on their protein levels by western blotting.
Results
CRISPR/Cas9, a genome engineering technique successfully knocked-out TMEPAI in the Hs578T TNBC cell line. Sequencing shows a frameshift mutation in TMEPAI. Western blot shows the absence of TMEPAI band on Hs578T KO cells.
Conclusion
TMEPAI gene was deleted in the TNBC cell line using the genomic editing technique CRISPR/Cas9. The deletion was confirmed by genome and protein analysis.
Keywords
CRISPR/Cas9, gene editing, knock-out cell lines
Genome editing methods have been developing recently. There are several genome editing technologies, namely zinc-finger nucleases (ZFNs), transcription activator-like effector nucleases (TALENs) and the clustered regularly interspaced short palindromic repeats/CRISPR-associated 9 (CRISPR/Cas9) system. ZFNs and TALENS have been put to practical use first. Both contain tethered endonuclease catalytic domains and deoxyribose nucleic acid (DNA) binding domain for introducing targeted DNA-double stranded break (DSB) in specific locus.1 Unlike them, the CRISPR/Cas9 system uses guide ribonucleic acid (gRNA) to recruit Cas9, an endonuclease, for pairing with the specific DNA region.1,2 Compared to previous techniques, the CRISPR/Cas9 system is easily designed, and gives more efficient multi genes editing for variety of cells and organism.1 DSB is repaired by DNA repairing pathways, nonhomologous end joining (NHEJ) or homologydirected repair (HDR) and undergo genome editing.3 NHEJ frequently generates insertion or deletion mutation. HDR mediates DNA editing in the presence of exogenous donor DNA.4
Triple negative breast cancer (TNBC) is a leading breast cancer causing death although it is found only in 15–25% of total breast cancer patients. It has no targeted therapy due to the lack of expression of estrogen receptor (ER), progesterone receptor (PR) and human epidermal growth factor receptor-2 (HER2). TNBC tends to grow more rapidly, frequently occurs in young women, more frequently metastasizes to lungs and brain, and more readily acquires drug resistance, compared to non-TNBC.5,6 Therefore, understanding the molecular mechanism in pathogenesis to determine targeted therapy and drug resistance mechanism in TNBC is necessary.
TMEPAI was found highly expressed in 68.8% of TNBC patients.5 Kaplan Meier (KM) plot showed that high TMEPAI expression correlates with poor prognosis. Previous research proved TMEPAI as a direct target gene and acts as a negative regulator by regulating duration and intensity of Transforming growth factor- β (TGF-β) responses in TGF-β Smad-dependent pathway.7 Additionally, TMEPAI can degrade phosphatase and tensin homolog (PTEN) by recruiting E3 ubiquitin ligase through its PY motifs to activate phosphatidylinositol-3-kinase/a serine/threonine kinase (PI3K/AKT) pathway.5 Furthermore, knock-down of TMEPAI in lung cancer reduces in vivo the xenograft tumor forming ability and the sphere forming ability. Thus, TMEPAI is thought to be a possible oncogenic protein.8 However, the molecular mechanisms of TMEPAI in TNBC is not clear yet.
We aimed to apply CRISPR/Cas9 technique to knockout TMEPAI in the triple negative breast cancer cell line. Knocking-out cell lines will be useful for further experiments to elucidate the TMEPAI function on tumorigenesis and drug resistance.
METHODS
Cell culture
The TNBC cell line, Hs578T was obtained from the American type culture collection (ATCC). Hs578T were cultured in Dulbecco’s modified essential medium (DMEM, Invitrogen) which is supplemented with 10% fetal bovine serum (FBS, Gibco), 10 μg/mL insulin, and 100 units/ ml of penicillin G and 0.1 mg/ml of streptomycin sulfate (Wako). Cells were maintained in 5% CO2 incubator at 37°C.
Plasmid construction
The gRNA against for TMEPAI exon2 locus was designed in silico using the CRISPR design online tool.9 The oligonucleotide primers have a default sequence for forward TTTCTTGGCTTTATATATCTTGTGGAAAGGACGAAACACCGtgcacggtccttcatcagc and reverse GACTAGCCTTATTTTAACTTGCTATTT CTAGCTCTAAAACgctgatgaaggaccgtgcaC.
Forward and reverse oligonucleotide were annealed by the standard method and followed by extension using PrimeSTAR DNA polymerase (Takara Bio). Extended 100 bps of doublestrand DNA was purified using polymerase chain reaction (PCR) purification kit (Qiagen) and inserted into gRNA cloning vector (Addgene plasmid #41824), which was a gift from George Church. It amplified in E. Coli and purified them using a miniprep purification kit (Qiagen). It was followed by sequencing before use.
Transfection
Hs578T cells were transfected with plasmid DNA using the polyethylenimine (PEI) transfection reagent. 1x105 cells were seeded on a 6-wells microplate one day before transfection. 1.5 μg plasmid DNA (500 ng gRNA, 500 ng human-Cas9, and a 500 ng puromycin resistant gene) in 4.5μL PEI, and incubate for 30 minutes. Medium was changed four to six hours after transfection.
Western blotting
Cells were collected with Tris buffer (62.5 mM Tris-HCl pH 7.5) and lysed using sodium dodecyl sulfate (SDS)-sample buffer (4% SDS, 1.44 M β-mercaptoethanol, 20% glycerol, 125 mN Tris pH 7.4, 0.002% bromophenol blue). Cell lysates were subjected to sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE). Proteins were then electro-transferred to a mix Polyvinylidene difluoride (PVDF) membrane (GE Healthcare) and subjected to immunoblot analysis. The primary antibodies used were rabbit anti-TMEPAI (homemade polyclonal antibody, 1:250)8 and mouse anti-β-actin antibody (Sigma, 1:5,000). Reacted antibodies were detected with horseradish peroxidase-conjugated anti-mouse IgG (GE Healthcare, 1:10,000), anti-rabbit IgG (GE Healthcare, 1:10,000), and Immuno-Star Zeta (Wako). EZ-capture MG (ATTO) was used for the detection of chemiluminescence.
RESULTS
Hs578T cells to induce DSB on TMEPAI exon 2 gene locus
We transfected gRNA expression vector together with human-Cas9 and puromycin resistant gene expression plasmids into Hs578T cells to induce DSB on TMEPAI exon 2 gene locus (Figure 1A). Puromycin selection was done for 72 hours since 24-post transfection. Surviving cells were seeded on a 10-cm dish to isolate single clone (two to three weeks). Then we expanded the grown cells for isolating protein. We analyzed TMEPAI expression by DNA sequencing (Figure 1C) and western blot (Figure 2).
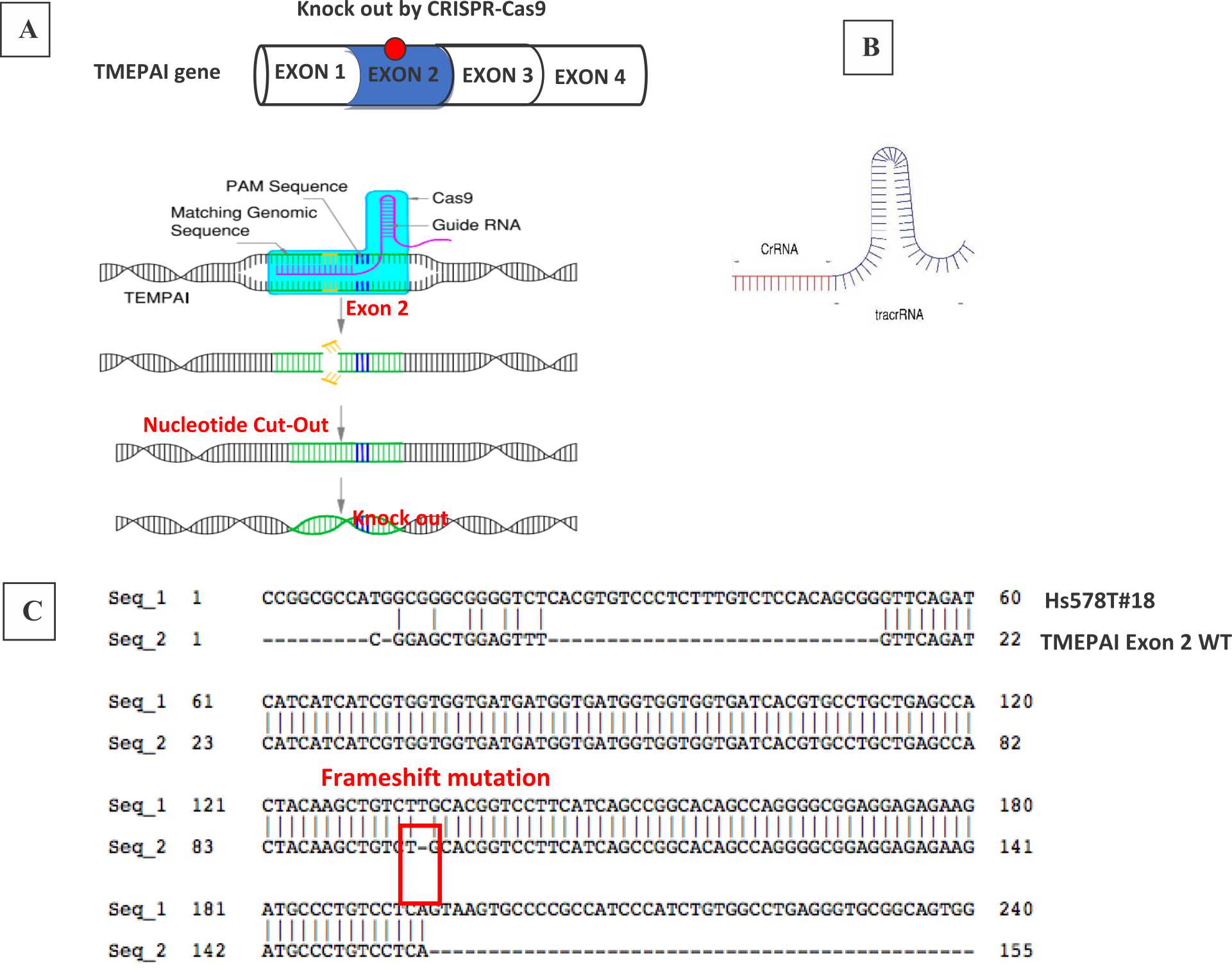
Figure 1. CRISPR-Cas9 system to knock-out TMEPAI. (A) CRISPR-Cas9 systems in Hs578T, the triple negative breast cancer cell line, (B) gRNA containing CrRNA which will direct Cas9 nuclease and tracrRNA which will recruit Cas9, (C) frameshift mutation due to knock-out the TMEPAI gene by CRISPR-Cas9 in Exon 2 TMEPAI.
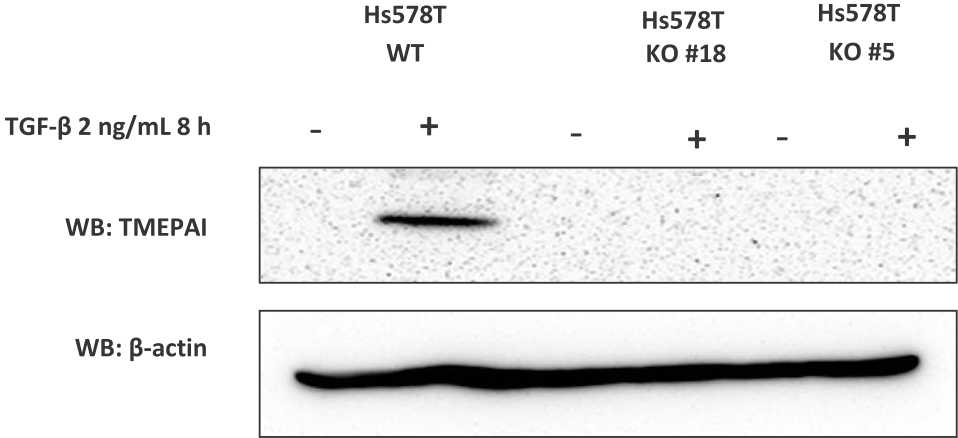
Figure 2. Establish knock-out TMEPAI in triple negative breast cancer cell line. The expression of TMEPAI protein in WT and KO Hs578T cells was measured by western blot analysis after TGF-β treatment. β-actin was used as a loading control
TNBC cell line, Hs578T responded to TGF-β stimulation and expressed TMEPAI eight hours after the treatment of TGF-β (2 ng/mL). However, KO-TMEPAI cells (clone #18 and #5) did not express TMEPAI protein (Figure 1C). The data indicate that we successfully established the KO-TMEPAI TNBC cell using the CRISPR-Cas9 system.
DISCUSSION
Previously, Cong et al11 and Mali et al12 demonstrated that Cas9 and gRNA can be used for genome editing in human cells. Instead of other techniques, it works as an advanced technique which is affordable, easy to use, and high-throughput. Neverthless similar to other techniques, it might make off-target cuts.13 The gRNA contains a targeting sequence (crRNA sequence) and a Cas9 nuclease-recruiting sequence (tracrRNA) (Figure 1B). The crRNA region (red color) is a nucleotide sequence that is homologous to a region in a TMEPAI sequence and the tracrRNA region directs Cas9 to the targeted sequence on genomic DNA. While tracrRNA interacts with Cas9, endonuclease cleaves the targeted sequence.1,2 Our results showed that crispr-cas9 successfully knocked out TMEPAI in exon 2. We got the edited genomic sequence in our cell line. This edited gene is inheritable to their daughter cells. In our study, the cells which were used in genomic and proteomic analyses came from passage No. 3-5.
Our gene of interest, TMEPAI, is also known as prostate transmembrane protein androgeninduced 1 (PMEPA1) and solid tumor-associated gene1 (STAG1). The TMEPAI gene encodes four protein isoforms, a, b, c, and d. Each has a distinct sequence in exon 1.14 Therefore, we could not target at exon 1. The sequence in exon 2 is commonly used for all isoforms. Hence, we targeted at exon 2 to edit all TMEPAI isoforms.
It was first found as an androgen-regulated gene which is related to a relapsed tumor in the human prostate cancer xenograft model that was strongly androgen-dependent.15 Currently, TMEPAI has been proven to enhance the tumorigenic activity in ovarian, breast and lung cancer cells,8,16 while in TNBC, TMEPAI promotes TGF-β dependent growth, motility and invasion.5 Taken together, TMEPAI is suspected as oncogenic protein. In contrast to above results, Fournier et al17 recently reported that low of TMEPAI correlates with poor metastasis-free survival and TMEPAI knockdown increases prostate cancer metastases in a mouse model.17 All the opposite results suggested that TMEPAI has an important function in cancer. However, how role of TMEPAI in cancer remained unclear. So, our cell line can be used in further experiments to elucidate the role of TMEPAI in TNBC pathogenesis to determine target therapy, drug resistance mechanism or other relevant research.
Instead of androgen, TMEPAI was induced by epidermal growth factor (EGF) in breast and ovarian cancer cell lines and in breast primary tumors,16 and TGF-β in many cancer cell lines.7,8,18 Singha et al5 reported that TGF-β is a stronger inducer of TMEPAI expression compared to EGF and lysophasphatidic acid (LPA). Notwithstanding, TMEPAI is the highest inducible gene of TGF-β, and TGF-β has a pivotal role in cancer. Therefore, we are concerned with TGF-β pathways to distinguish clearly between wild-type (WT) and knock-out (KO) cell lines of TMEPAI.
This study used one triple negative breast cancer cell line, Hs578T. Considering that TNBC is a diverse and heterogenous group of breast cancer type, we can not generalize this system in other triple negative breast cancer cell lines. Despite the limitation, our study clearly shows that our designed gRNA can edited the TMEPAI genome and further disperses TMEPAI protein expression. In conclusion, the TMEPAI gene was deleted in the TNBC cell line using the genomic editing technique CRISPR/Cas9, and the deletion was confirmed by genome and protein analyses. This TMEPAI knock-out cell line can be used for further experiments to elucidate the role of TMEPAI in TNBC pathogenesis to determine target therapy, drug resistance mechanism or other relevant research.
Conflicts of Interest
The authors affirm no conflict of interest in this study.
Acknowledgment
None.
REFERENCES
- Ran FA, Hsu PD, Wright J, Agarwala V, Scott DA, Zhang F. Genome engineering using the CRISPR-Cas9 system. Nat Protoc. 2013;8(11):2281–308.
- Tycko J, Myer VE, Hsu PD. Methods for optimizing CRISPR-Cas9 genome editing specificity. Mol Cell. 2016;63(3):356–70.
- Sánchez-Rivera FJ, Jacks T. Applications of CRISPRCas9 system in cancer biology. Nat Rev Cancer. 2015;15(7):387–95.
- Weinberg RA. The biology of cancer. 2nd edition. New York: Garland Sence; 2013.
- Singha PK, Pandeswara S, Geng H, Lan R, Venkatachalam MA, Saikumar P. TGF-β induced TMEPAI/PMEPA1 inhibits canonical Smad signaling through R-Smad sequestration and promotes non-canonical PI3K/Akt signaling by reducing PTEN in triple negative breast cancer. Genes Cancer. 2014;5(9-10):320–36.
- Foulkes WD, Smith IE, Reis-Fielho JS. Triple negative breast cancer. N Eng J Med. 2010;363:1938–48.
- Watanabe Y, Itoh S, Goto T, Ohnishi E, Inamitsu M, Itoh F, et al. TMEPAI, a transmembrane TGF-β- inducible protein, sequesters Smad proteins from active participation in TGF-beta signaling. Mol Cell. 2010;37(1):123–34.
- Vo Nguyen TT, Watanabe Y, Shiba A, Noguchi M, Itoh S, Kato M. TMEPAI/PMEPA1 enhances tumorigenic activities in lung cancer cells. Cancer Sci. 2014;105(3):334–41.
- CRISPRdirect-Rational design of CRISPR/Cas target [available from: https://crispr.dbcls.jp]
- Itoh S, Thorikay M, Kowanetz M, Moustakas A, Itoh F, Heldin CH, et al. Elucidation of Smad requirement in transforming growth factor-beta type I receptorinduced responses. J Biol Chem. 2003;278(6):3751–61.
- Cong L, Ran FA, Cox D, Lin S, Barretto R, Habib N, et al. Multiplex genome engineering using CRISPR/Cas systems. Science. 2013;339(6121):819–23.
- Mali P, Yang L, Esvelt KM, Aach J, Guell M, DiCarlo JE, et al. RNA-guided human genome engineering via Cas9. Science. 2013;339(6121):823–6.
- Falahi F, Sgro A, Blancafort P. Epigenome engeenering in cancer: fairytale or a realistic path to the clinics?. Front Oncol. 2015;5:1–11.
- Vo Nguyen TT. Tumorigenic function of TMEPAI in cancer. Tulips University of Tsukuba Library. 2014. p.13
- Xu LL, Shanmugam N, Segawa T, Sesterhenn IA, McLeod DG, Moul JW, et al. A novel androgen- regulated gene, PMEPA1, located on chromosome 20q13 exhibits high level expression in prostate. Genomic. 2000;66(3):257–63.
- Giannini G, Ambrosini MI, Di Marcotullio L, Cerignoli F, Zani M, MacKay AR, et al. EGF- and cell-cycle-regulated STAG1/PMEPA1/ERG1.2 belongs to a conserved gene family and is overexpressed and amplified in breast and ovarian cancer. Mol Carcinog. 2003;38(4):188–200.
- Fournier PG, Juárez P, Jiang G, Clines GA, Niewolna M, Kim HS, et al. The TGF-β signaling regulator PMEPA1 supresses prostate cancer metastases to bone. Cancer Cell. 2015;27(6):809–21.
- N, Triaspolitica. "Kanker Payudara: Informasi, Penyebab, Gejala, Stadium Dan Pengobatan." Mau Nanya Dong Dok. N.p, 28 June 2017. Web. 30 June 2017. <https://nanyadongdok.blogspot.com/2017/06/kankerpayudara-informasi-penyebab.html>.
Copyright @ 2017 Authors. This is an open access article distributed under the terms of the Creative Commons Attribution-NonCommercial 4.0 International License (http://creativecommons.org/licenses/by-nc/4.0/), which permits unrestricted non-commercial use, distribution, and reproduction in any medium, provided the original author and source are properly cited.
mji.ui.ac.id